Stata Tips #18 - Fitting more complex Bayesian models with BUGS, Stan or JAGS – from inside Stata
In this post, we will look at a couple of community-contributed commands for fitting Bayesian models to your data. Stata 14 introduced Bayesian functionality for the first time with bayesmh, and version 15 took this further with the bayes: prefix, which can conveniently be added before calling any of 45 estimation commands (just as you might type bootstrap: or svy:), but you can also fit bespoke models with external, free software: BUGS, Stan and JAGS.
Why would I need anything other than Stata's Bayesian commands?
BUGS, Stan and JAGS have been around since before Stata's bayesmh and bayes:. They used to be the only way to fit these kinds of models, so you might well find that someone has written code for one of them that you can easily adapt to your needs.
In particular, The BUGS Book and the Stan manual and case studies contain many examples of specialised models, and wise advice on adapting it.
Also, there are some kinds of model that Stata can't fit. At the time of writing, this includes imputing of missing or censored data inside a Bayesian model, Gaussian processes, or geospatial models.
Installing commands for BUGS and JAGS
Various commands for interfacing between Stata and WinBUGS, OpenBUGS and JAGS were written by John Thompson of the University of Leicester in the UK. You can download them all from Stata Press, because they accompany his book Bayesian Analysis With Stata:
net from http://www.stata-press.com/data/bas
net get bayes
do bayes
Then, follow the instructions in the readme.txt file to complete installation. You will need the WinBUGS folder location, so read on if you haven't already installed it.
To work with JAGS, you will need the experimental v2 versions of these commands. We won't go into them in this post, so there's no need to install them now, but I do suggest you consider JAGS in preference to BUGS, as a more actively-maintained and used piece of software, and one that works on all operating systems.
If you are familiar with the version control software, Git, you might prefer to clone my GitHub repository to get all the files in one place, as well as the experimental v2 versions, which we'll discuss below: git clone https://github.com/robertgrant/winbugsfromstatav2 If you don't do git, you can just point your browser to https://github.com/robertgrant/winbugsfromstatav2/archive/master.zip and unzip this into your personal ado path.
(What's my personal ado path? It's a folder on your computer where your own Stata commands get added, like stuff you install from SSC. You can find it by typing in Stata: sysdir, then looking for the PERSONAL directory.

This package was originally aimed at running WinBUGS, a graphical user interface to the BUGS program that the Medical Research Council Biostatistics Unit in Cambridge created for Windows computers, and some years ago stopped actively developing. You will need to install WinBUGS if you want to use it (there are instructions here). If you are not using Windows, or just want to use OpenBUGS or JAGS (which both run the same algorithm, the Gibbs sampler, but unlike WinBUGS, do not have a graphical user interface), then you will need to work with John's experimental v2 commands. They are included in my GitHub repository.
You will find instructions to install OpenBUGS on Windows or Linux (there is no guarantee that the Linux version will behave as expected on Mac) here and JAGS here. My personal advice is to use JAGS, which is actively maintained and has a large pool of expert users, who are likely to have detected and fixed any problems before you encounter them. JAGS is the only option for Mac anyway.
If you are considering using JAGS, check out John's blog post on the (relatively minor) differences. Once you install JAGS, look in the Readme.rtf file that is installed to find the location of the executable program on your operating system. You will need to tell Stata this location.
Installing commands for Stan
To use Stan, you need to install CmdStan, and there are instructions here, particularly in Appendix B of the CmdStan manual. Note that, for Windows, you should install RTools as well, and at the time of writing, it may be best to avoid CmdStan 2.18.0, which has a lot of speed improvements but many Windows users have had problems installing it.
Like JAGS, you need to know where it has been installed on your computer, so that you can tell Stata when you run it.
Then, installing the Stata commands is simple because they are on SSC:
ssc install stan
And, if you are using Windows, you will need the windowsmonitor command:
ssc install windowsmonitor
Running a simple model in WinBUGS
Now, having done the tiresome business of installation, we can start sending data and model specification from Stata to some of these Bayesian programs. We will use the simple regression from my previous blog post on Bayes in Stata:
sysuse auto, clear
regress price mpg
We could just put the bayes: prefix in from of this regress command, but the default priors are not suitable here, so to get better results, we specified a normally-distributed prior for the slope, and uniformly-distributed priors for the constant term and the residual variance:
bayes, prior({price: mpg}, normal(0, 4000000)) ///
prior({price: _cons}, uniform(-10000,25000)) ///
prior({sigma2}, uniform(0.1, 5000)) ///
initial({price: _cons} 12000 {price: mpg} -300 {sigma2} 500) ///
: regress price mpg
If we wrote out this model algebraically, it would be something like this:
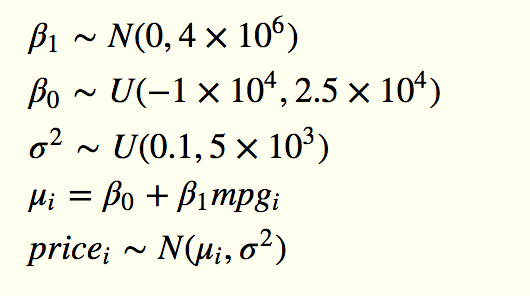
The first three lines are the priors, the fourth deterministically sets the linear predictor mu, given values drawn from the priors, and the fifth defines the likelihood. In BUGS, JAGS or Stan, you specify the model in individual statements like this.
The model code for BUGS or JAGS is the same:
model {
beta0 ~ dunif(-10000,25000)
beta1 ~ dnorm(0,0.00000025)
tau ~ dgamma(0.001,0.001)
for (i in 1:74) {
mu[i] <- beta0 + (beta1 * mpg[i])
price[i] ~ dnorm(mu[i],tau)
}
}
You might notice two things that differ from the Stata 15 specification: instead of variance, we have to specify precision (1/variance), and BUGS is much less tolerant of priors on precisions other than the gamma distribution, so here we are using a diffuse gamma. (Note that this has been heavily criticised...)
There are some ways of specifying this model inside the do-file, but to keep it simple, you can just type it up inside WinBUGS or a text editor (like Notepad, Atom, Sublime Text, vim, or whatever). Here, I saved it as autommodel.txt. The rest of the files that WinBUGS will need are generated for you by John Thompson's excellent commands.
wbslist or wbsarray can be used to write your data out into a text format. In my experience, you need to use the format option to get readable results. wbslist will also write out the initial values. Here, we are just running one chain.
wbsscript writes the script file for WinBUGS that tells it where to find the model, the initial values and data, how many iterations to run, and so on. If you know MCMC analysis, the options are pretty self-explanatory. The noquit option keeps WinBUGS open, and Stata won't proceed until you shut it, which is handy while you are debugging and getting your code right.
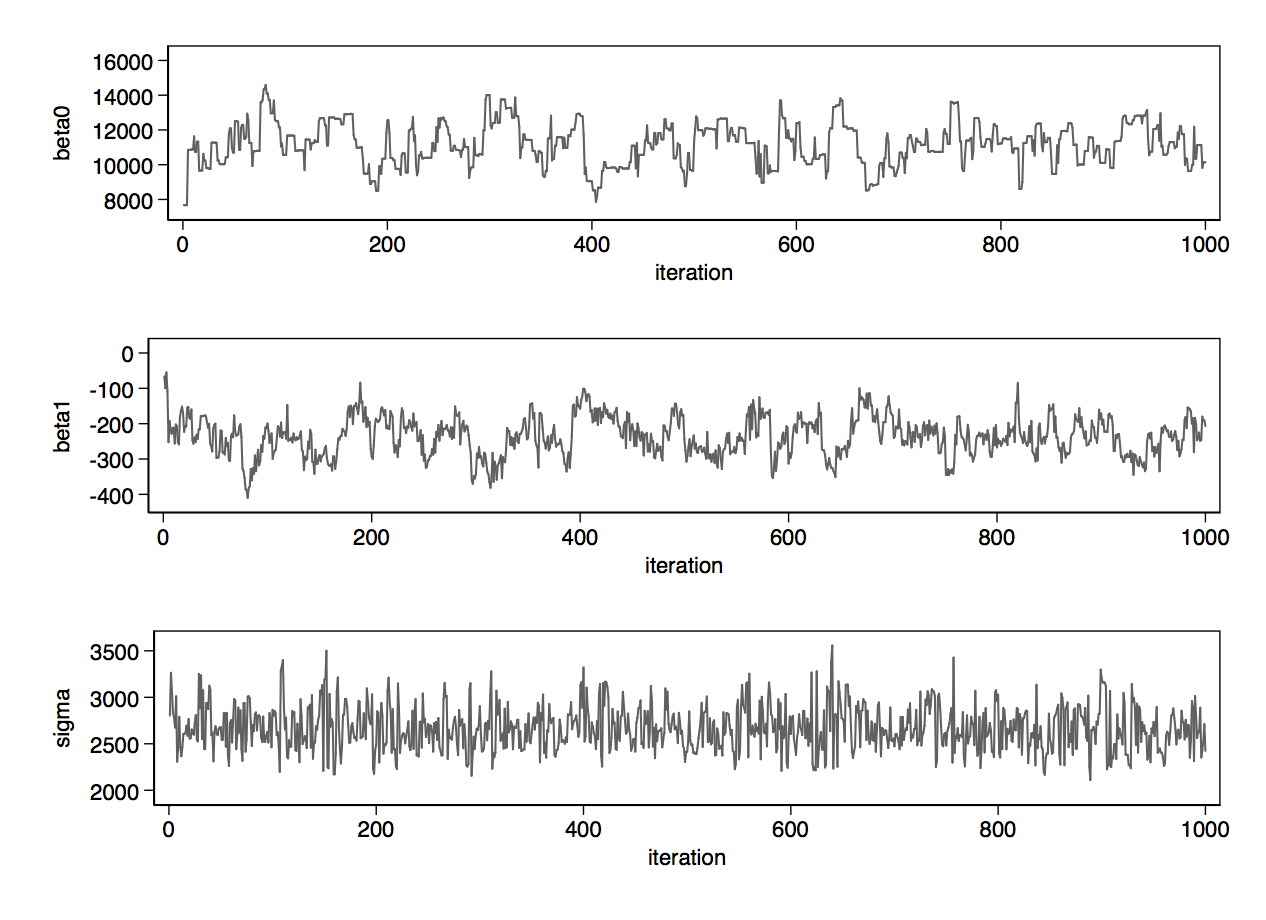
Having obtained your results by running wbsrun, wbscoda reads the iterations of the Markov chains into Stata, which you can summarise statistically with mcmcstats and visually with mcmctrace, as well as other approaches.
Here are the outputs from mcmcstats:
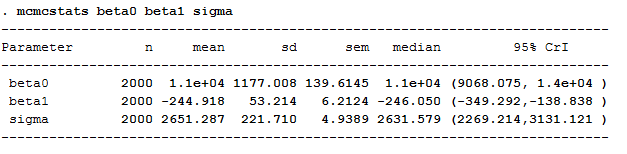

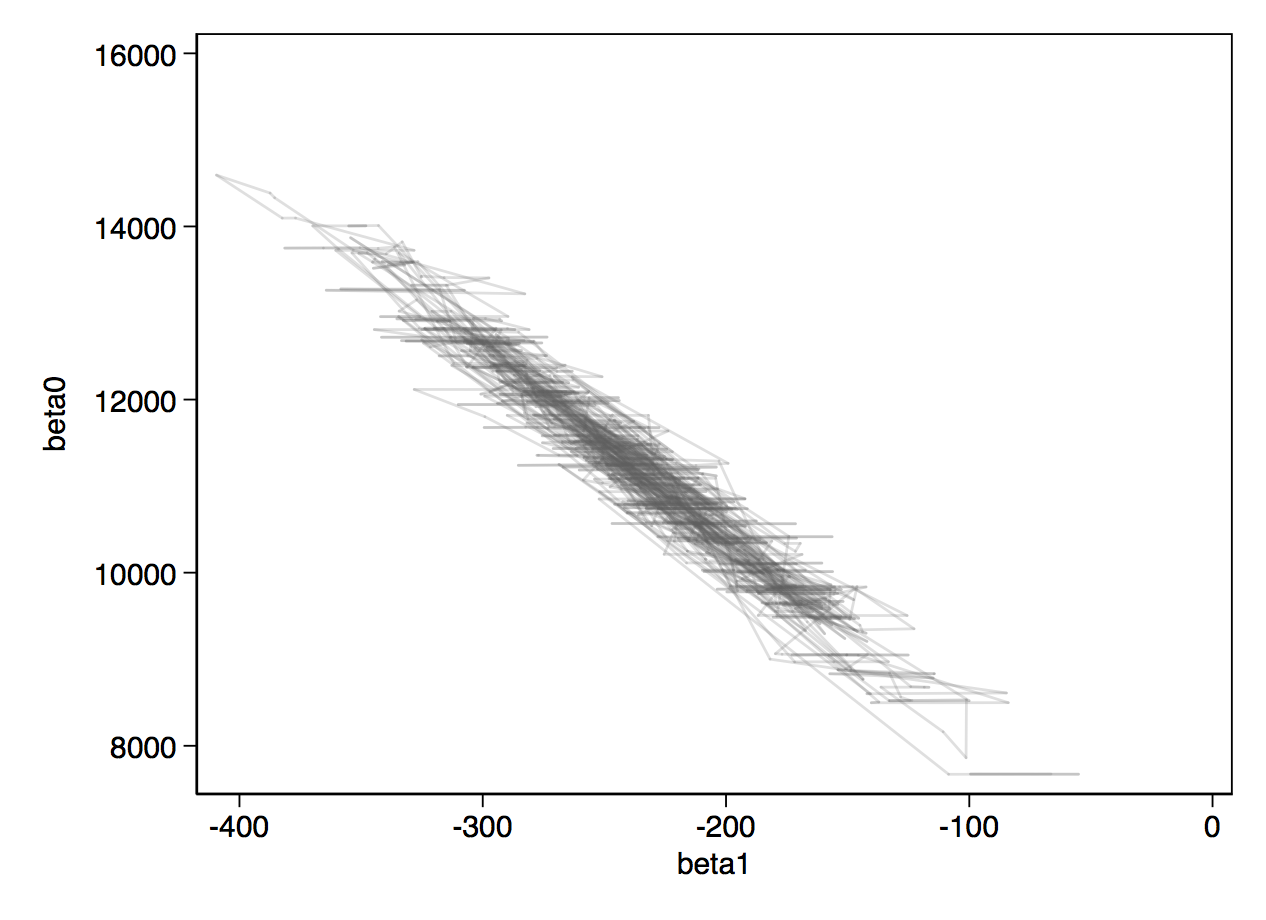
Below, you can see the traceplots from mcmctrace, then a bivariate trace where we simply get a line chart of the two betas, which we expect to be quite strongly negatively correlated.
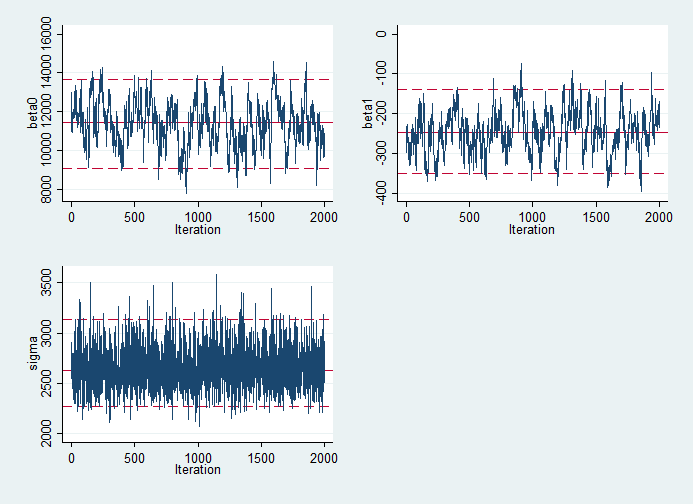
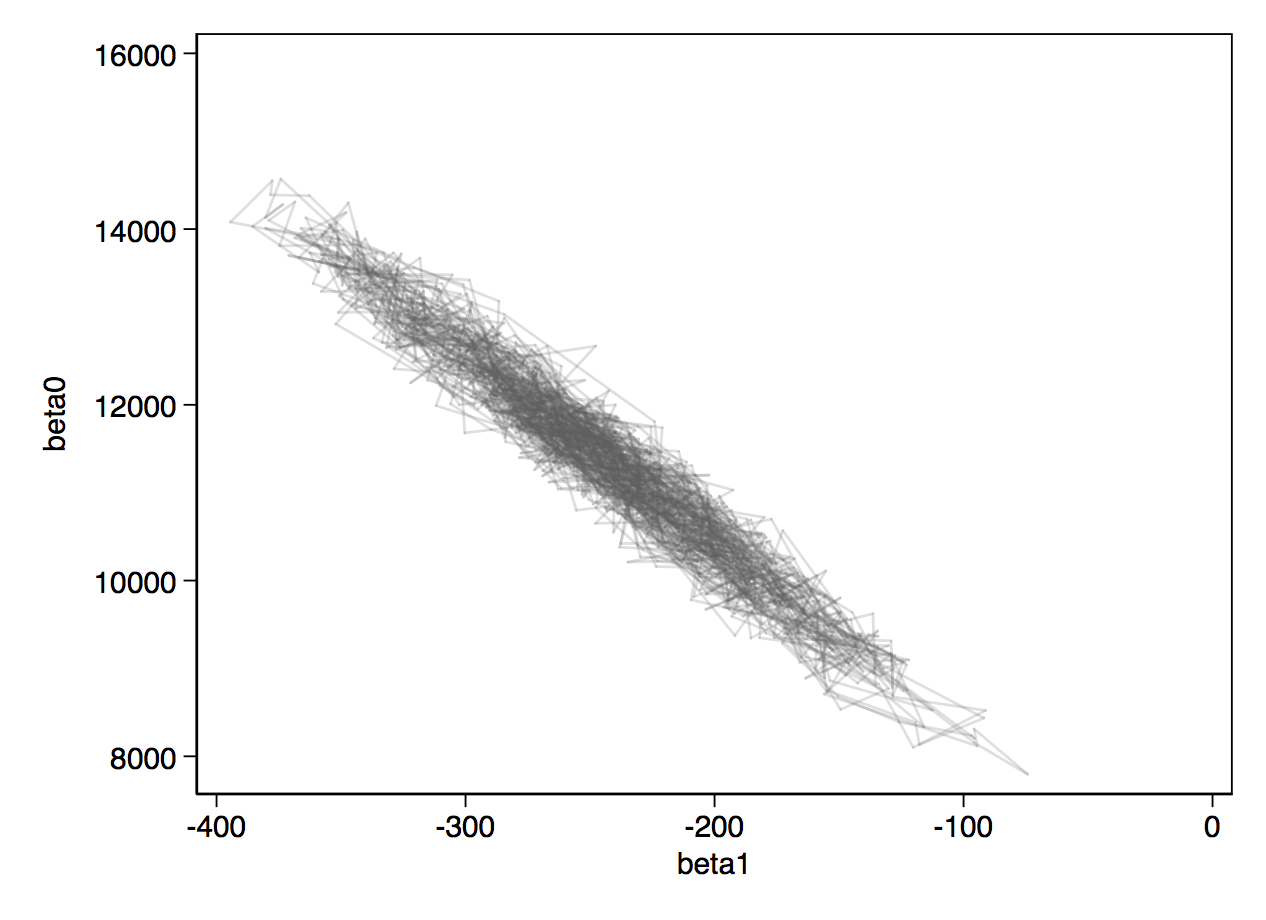
If you wanted to use OpenBUGS instead, you can just add the openbugs option to your wbsscript, wbsrun and wbscoda commands. Thompson's v2 versions of the commands are an experimental approach to extending this to JAGS. They have not been fully tested so you should use them with caution.
Running a simple model in Stan
To do the same analysis in Stan, we need a somewhat different model file:
data {
int N;
real mpg[N];
real price[N];
}
parameters {
real beta0;
real beta1;
real<lower=0> sigma;
}
model {
real mu[N];
beta0 ~ uniform(-10000,25000);
beta1 ~ normal(0,2000);
sigma ~ uniform(0.1,5000);
for(i in 1:N) {
mu[i] = beta0 + (beta1*mpg[i]);
price[i] ~ normal(mu[i],sigma);
}
}
Data and parameters have to be declared before they can be used; we assign them to be integers or real numbers in this example. This helps the computer work more efficiently. Also, instead of precision, we can specify the standard deviation, which is usually more intuitive. The algorithm used by Stan is stable with a wide range of prior distributions, so we can use a uniform on the standard deviation, and we do not have to specify intial values (usually). This model file is saved as auto.stan. Then, we store the number of rows as a global N, and use the stan command:
count
global N=r(N)
stan mpg price, globals("N") modelfile("auto.stan") ///
cmdstandir("/Users/robert/cmdstan-2.16.0") load
Here are the outputs:
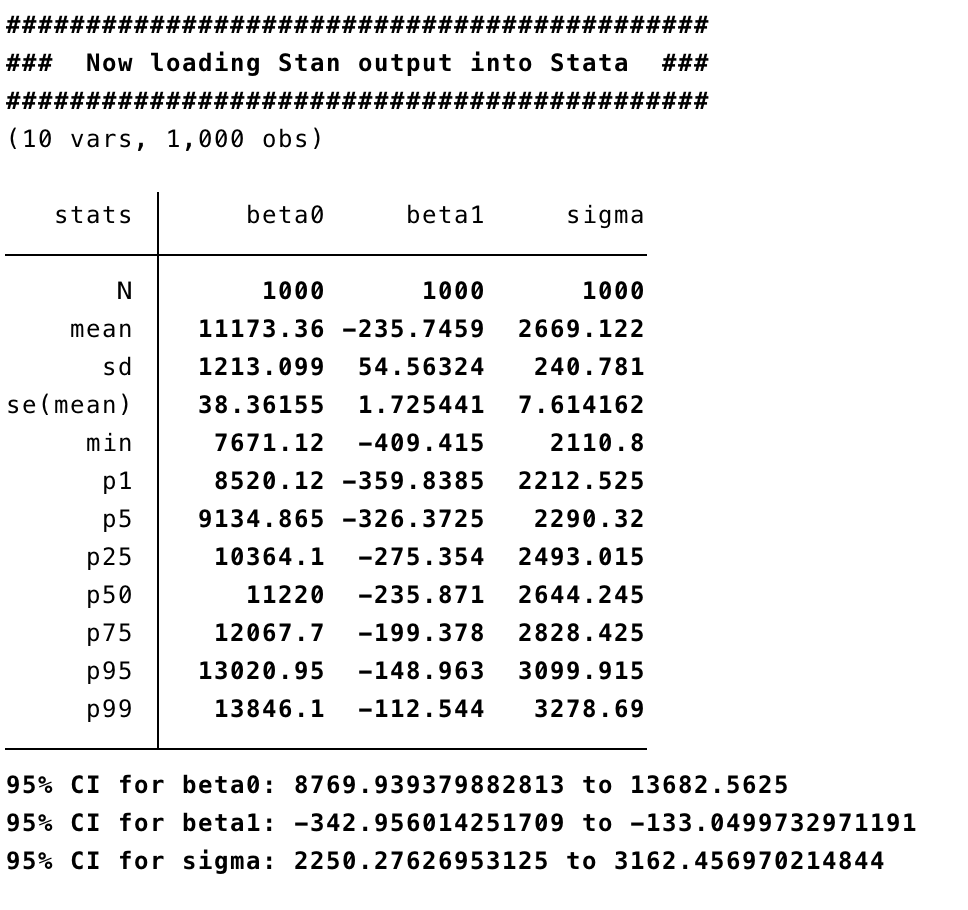
For a very simple example with only 74 observations and 3 parameters, Stan takes notably longer than BUGS. As the size of the problem grows, or as parameters become more correlated, or as the priors become more bespoke (non-conjugate), Stan will win on speed. The load option reads the chain back into Stata, and then we can generate traceplots and a bivariate trace on the betas.
The do-file for all the commands (except installations) in this post can be downloaded by clicking here.